B7 - Role of Trib1 in obesity and metabolism - Tissue-specific effects and target molecules
We had previously identified the gene Trib1 (encoding Tribbles homolog-1) as a novel regulator of lipoprotein metabolism in mice. Initial data from the literature and our group suggested that Trib1 might fulfill a broader function in the control of metabolism. During the first funding period we established that Trib1 affects glycemic control, insulin resistance, adipose tissue function and overall susceptibility to high-fat diet-induced obesity.
Our findings suggest that Trib1 is involved in the regulation of multiple metabolic pathways. To reduce the complexity of the phenotype observed in whole-body Trib1 knockout mice, we now aim to delineate tissue-specific effects and contributions of Trib1. Metabolic and functional characterization of these models will elucidate how Trib1-depletion influences insulin sensitivity, obesity associated adipose tissue dysfunction and ectopic fat deposition in relevant tissues.
In a second related aim, we propose to follow-up promising candidate genes/molecules that are likely to be important in mediating the phenotype of Trib1-/- mice. We will capitalize on data obtained during the first funding period, where we identified several regulators of lipid metabolism as promising candidates. In addition to their role in metabolism, these factors have been described to modulate cellular stress and tissue inflammation, important mechanisms contributing to adipose tissue dysfunction. However, their role in adipose tissue biology has not been studied in detail and is not entirely understood. We will use samples from the human adipose tissue biobank (Project B1), well-phenotyped lean and obese participants from cross-sectional (Leipzig LIFE studies) and different weight-loss intervention cohorts (e.g. CENTRAL RCT, project B8), combined with mechanistic studies in tissue culture and appropriate mouse models, to unravel the function of these candidate molecules in the pathogenesis of obesity.
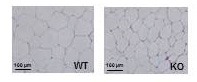
Figure 1. Hematoxylin eosin staining of white adipose tissue from male KO and Trib1-WT mice after 16 weeks of high-fat feeding.
Ecker J, Benedetti E, Kindt ASD, Höring M, Perl M, Machmüller AC, Sichler A, Plagge J, Wang Y, Zeissig S, Shevchenko A, Burkhardt R, Krumsiek J, Liebisch G, Janssen KP. The colorectal cancer lipidome - identification of a robust tumor-specific lipid species signature. Gastroenterology. 2021 May 14:S0016-5085(21)02979-6.
Krautbauer S, Blazquez R, Liebisch G, Hoering M, Neubert P, Pukrop T, Burkhardt R, Sigruener A. Application of Lipid Class Ratios for Sample Stability Monitoring-Evaluation of Murine Tissue Homogenates and SDS as a Stabilizer. Metabolites. 2021 Apr 27;11(5):277.
Höring M, Ejsing CS, Krautbauer S, Ertl VM, Burkhardt R, Liebisch G. Accurate quantification of lipid species affected by isobaric overlap in Fourier-Transform mass spectrometry. J Lipid Res. 2021 Feb 15:100050.
Orsó E, Burkhardt R. ATP-citrate lyase: a driver of metabolism and histone acetylation. Curr Opin Lipidol. 2020 Dec;31(6):362-363.
Blucher C, Iberl S, Schwagarus N, Muller S, Liebisch G, Horing M, Hidrobo MS, Ecker J, Spindler N, Dietrich A, Burkhardt R, Stadler SC.Secreted factors from adipose tissue reprogram tumor lipid metabolism and induce motility by modulating PPARα/ANGPTL4 and FAK. Mol Cancer Res. 2020 Aug 28:molcanres.1223.2019.
Höring M, Ekroos K, Baker PRS, Connell L, Stadler SC, Burkhardt R, Liebisch G. Correction of isobaric overlap resulting from sodiated ions in lipidomics. Anal Chem. 2020 Jul 16.
Jacobi T, Massier L, Klöting N, Horn K, Schuch A, Ahnert P, Engel C, Löffler M, Burkhardt R, Thiery J, Tönjes A, Stumvoll M, Blüher M, Doxiadis I, Scholz M, Kovacs P. HLA Class II Allele Analyses Implicate Common Genetic Components in Type 1 and Non–Insulin-Treated Type 2 Diabetes. J Clin Endocrinol Metab. 2020 Mar 1;105(3).
Blücher C, Zilberfain C, Venus T, Spindler N, Dietrich A, Burkhardt R, Stadler SC, Estrela-Lopis I. Single cell study of adipose tissue mediated lipid droplet formation and biochemical alterations in breast cancer cells. Analyst. 2019 Aug 13
Wagner R, Dittrich J, Thiery J, Ceglarek U, Burkhardt R. Simultaneous LC-MS/MS quantification of eight apolipoproteins in normal and hypercholesterolemic mouse plasma. J Lipid Res. 2019 Feb 5. pii: jlr.D084301
Dittrich J, Beutner F, Teren A, Thiery J, Burkhardt R, Scholz M, Ceglarek U. Plasma levels of apolipoproteins C-III, A-IV, and E are independently associated with stable atherosclerotic cardiovascular disease. Atherosclerosis. 2018 Nov 9;281:17-24.
Lalem T, Zhang L, Scholz M, Burkhardt R, Saccheti V, Teren A, Thiery J, Devaux Y; Cardiolinc™ network (www.cardiolinc.org). Cyclin dependent kinase inhibitor 1 C is a female-specific marker of left ventricular function after acute myocardial infarction. Int J Cardiol. 2018 Jul 10. pii: S0167-5273(18)32556-7.
Arndt L, Dokas J, Gericke M, Kutzner CE, Müller S, Jeromin F, Thiery J, Burkhardt R. Tribbles homolog 1 deficiency modulates function and polarization of murine bone marrow-derived macrophages. J Biol Chem. 2018 Jun 13. pii: jbc.RA117.000703.
Nickel A, Blücher C, Kadri OA, Schwagarus N, Müller S, Schaab M, Thiery J, Burkhardt R, Stadler SC. Adipocytes induce distinct gene expression profiles in mammary tumor cells and enhance inflammatory signaling in invasive breast cancer cells. Sci Rep. 2018 Jun 21;8(1):9482.
Pott J, Schlegel V, Teren A, Horn K, Kirsten H, Bluecher C, Kratzsch J, Loeffler M, Thiery J, Burkhardt R, Scholz M.Genetic Regulation of PCSK9 (Proprotein Convertase Subtilisin/Kexin Type 9) Plasma Levels and Its Impact on Atherosclerotic Vascular Disease Phenotypes. Circ Genom Precis Med. 2018 May;11(5):e001992.
Arndt L, Burkhardt R. Lipid metabolism: spotlight on modulators of lipoprotein lipase activity. Curr Opin Lipidol. 2018 Apr;29(2):164-165
Buerger F, Müller S, Ney N, Weiner J, Heiker JT, Kallendrusch S, Kovacs P, Schleinitz D, Thiery J, Stadler SC, Burkhardt R. Depletion of Jmjd1c impairs adipogenesis in murine 3T3-L1 cells. Biochim Biophys Acta. 2017;1863:1709-17.
Vausort M, Salgado-Somoza A, Zhang L, Leszek P, Scholz M, Teren A,Burkhardt R,Thiery J, Wagner DR, Devaux Y. Myocardial infarction-associated circular RNA predicting left ventricular dysfunction. J Am Coll Cardiol. 2016;68:1247-8.
Burkhardt R, Kirsten H, Beutner F, Holdt LM, Gross A, Teren A, Tönjes A, Becker S, Krohn K, Kovacs P, Stumvoll M, Teupser D, Thiery J, Ceglarek U, Scholz M. Integration of genome-wide SNP data and gene-expression profiles reveals six novel loci and regulatory mechanisms for amino acids and acylcarnitines in whole blood. PLoS Genet. 2015;11:e1005510.
Keller M, Schleinitz D, Förster J, Tönjes A, Böttcher Y, Fischer-Rosinsky A, Breitfeld J, Weidle K, Rayner NW, Burkhardt R, Enigk B, Müller I, Halbritter J, Koriath M, Pfeiffer A, Krohn K, Groop L, Spranger J, Stumvoll M, Kovacs P. THOC5 : a novel gene involved in HDL-cholesterol metabolism. J Lipid Res. 2013;54:3170-6.
Burkhardt R, Sündermann S, Ludwig D, Ceglarek U, Holdt LM, Thiery J, Teupser D. Cosegregation of aortic root atherosclerosis and intermediate lipid phenotypes on chromosomes 2 and 8 in an intercross of C57BL/6 and BALBc/ByJ low-density lipoprotein receptor-/- mice. Arterioscler Thromb Vasc Biol. 2011;31:775-84.
Burkhardt R, Toh SA, Lagor WR, Birkeland A, Levin M, Li X, Robblee M, Fedorov VD, Yamamoto M, Satoh T, Akira S, Kathiresan S, Breslow JL, Rader DJ. Trib1 is a lipid- and myocardial infarction-associated gene that regulates hepatic lipogenesis and VLDL production in mice. J Clin Invest. 2010;120:4410-4.
Lowe JK, Maller JB, Pe'er I, Neale BM, Salit J, Kenny EE, Shea JL, Burkhardt R, Smith JG, Ji W, Noel M, Foo JN, Blundell ML, Skilling V, Garcia L, Sullivan ML, Lee HE, Labek A, Ferdowsian H, Auerbach SB, Lifton RP, Newton-Cheh C, Breslow JL, Stoffel M, Daly MJ, Altshuler DM, Friedman JM. Genome-wide association studies in an isolated founder population from the Pacific Island of Kosrae. PLoS Genet. 2009;5:e1000365.
Burkhardt R, Kenny EE, Lowe JK, Birkeland A, Josowitz R, Noel M, Salit J, Maller JB, Pe'er I, Daly MJ, Altshuler D, Stoffel M, Friedman JM, Breslow JL. Common SNPs in HMGCR in micronesians and whites associated with LDL-cholesterol levels affect alternative splicing of exon13. Arterioscler Thromb Vasc Biol. 2008;28:2078-84.
Teupser D, Kretzschmar D, Tennert C, Burkhardt R, Wilfert W, Fengler D, Naumann R, Sippel AE, Thiery J. Effect of macrophage overexpression of murine liver X receptor-alpha (LXR-alpha) on atherosclerosis in LDL-receptor deficient mice. Arterioscler Thromb Vasc Biol. 2008;28:2009-15.